Last month Kevin talked about the “bipolar” lives of paleoanthropologists. This month I thought I would bring up another great attribute of working in this field, namely having the good fortune to be able to work with scientists across different labs with similar research interests. One of the best aspects about being a student in the HomPal PhD program at GWU is that the faculty members collaborate with researchers across the globe, which affords the students an opportunity to train in some amazing locations. Some of the places my fellow students have worked are South Africa, Kenya, Indonesia, Germany, and Finland, just to name a few. But, I went someplace even more exciting to me personally… Detroit.
I came to the HomPal program to study phylogenetics with Dr. Bernard Wood and I am especially interested in examining genotypic and phenotypic evolution of complex character traits. It is a really exciting time in phylogenetics because recent advances in DNA sequencing and bioinformatic software have allowed for an exponential increase in genomic data collection and have provided a deeper understanding of the evolutionary molecular relationships between humans and our closest living relatives. This technical revolution has allowed for some very extraordinary research, such as the recent publication of the Neanderthal genome.
To gain experience with these new phylogenomic techniques, last summer I traveled to the Center for Molecular Medicine and Genetics at Wayne State University School of Medicine in Michigan to complete a lab rotation with Drs. Derek Wildman and Kirstin Sterner (now at the University of Oregon) because I was drawn to their work in comparative genomics. OK, the location wasn’t as exotic as those visited by some of my fellow classmates, but for me it was the perfect choice.
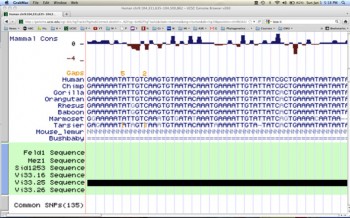
Since I am interested in examining the molecular evolution of the human phenotype using comparative genomic methods, my project focused on the molecular evolution of GRIN3A, a gene previously determined to play an important role in human brain plasticity. I learned phylogenomic and bioinformatic techniques necessary to investigate adaptive evolution in the regulatory and coding regions of GRIN3A in primate lineages in order to infer how selection has acted upon these elements.
When I came back to Washington D.C. in the fall I continued working on this project with Dr. Chet Sherwood at the GWU Laboratory for Evolutionary Neuroanatomy. He supplied me with tissue samples from 12 different primate species in order to analyze the gene expression of GRIN3A in the neocortex. He also put me in touch with the GW Institute for Neuroscience Biomarkers Analysis and Discovery Core where we extracted the RNA and are currently analyzing it with qPCR. Since we are looking at the mRNA, we must first use reverse transcriptase to convert mRNA into cDNA and then amplify it with qPCR.
Not every program can offer its students such opportunities. It has been a fulfilling project and I am lucky to be working with the researchers in these three very different labs.